Protein acylation refers to all modifications that covalently bind acyl groups to a protein. Acylation occurs in all living organisms and is carried out by specific acyl transferases. The structure and size of the acyl groups varies greatly, from small moieties such as acetate to long chain fatty acids such as palmitic acid. The moieties have to be activated first before they are attached to different amino side chain residues1. The most common form of acylation is acetylation. However, this post will be dedicated to two other important and frequently occurring acylations called N-myristoylation and S-palmitoylation2.
N-Myristoylation
Overview
N-Myristoylation refers to the covalent linkage of a myristic acid to the N-terminal glycine via an amide bond of many eukaryotic and viral proteins. The reaction is catalyzed by N-myristoyltransferase, is irreversible and affects the hydrophobicity of the protein.
| pKa | NC | Loss | Gain | Deltamass | H | AA | UV-Spec | Pattern |
| – | No | H | C14H27O | Av: 210.3562 M: 210.1984 | – | ^ | – | ^(?=G) |
In-depth mechanism
N-Myristoylation describes the process of covalently attaching a myristate moiety, a 14-carbon saturated fatty acid, onto an N-terminal glycine via amide bond. The myristate moiety is activated by coenzyme A (CoA), enabling the N-myristoyltransferase (NMT) that catalyses the reaction to convert the substrate3–6. The mechnanism of N-myristoylation is shown in figure 1.
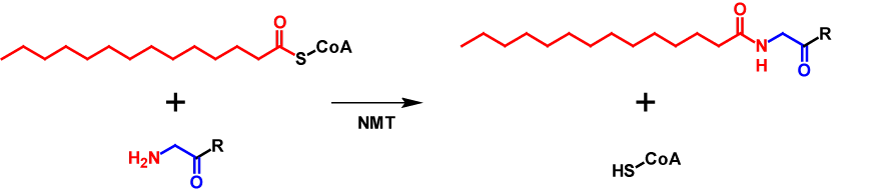
Roughly 0.5 % of all eukaryotic proteins are N-myristoylated. For proteins to be N-myristoylated, the first to amino acids need to be a methionine followed by a gylcine. Often a serine or threonine is present at position 5. N-Myristoylation is typically a co-translational modification, but in a few cases also post-translational. First, methionine aminopeptidase removes the initiator methione and then the myristate moiety is attached to the N-terminal glycine3,6. This modification is generally irreversible4 and leads to an increase of the molecular mass by 210 Da7. Since myristate is a hydrophobic molecule, it affects intracellular localisation and functions of the modified proteins. These include influencing protein–protein interactions, enhancing interactions of the protein with organelle or plasma membranes and affects protein stability5.
S-Palmitoylation
Overview
S-Palmitoylation is a reversibel post-translational modifaction, whereby a palmitic acid is covalently attached to cysteine residue of a protein. The reaction is catalyzed by protein palmitoyl acyltransferases. This modification strongly alters the hydrophobicity of the protein.
| pKa | NC | Loss | Gain | Deltamass | H | AA | UV-Spec | Pattern |
| – | No | H | C16H31O | Av: 238.4094 M: 238.2297 | – | C | – | – |
In-depth mechanism
S-Palmitoylated proteins contain covalently attached palmitate moieties to one or more cysteine residues. Palmitate is a 16-carbon long saturated fatty acid, which is also activated by binding to CoA3,8,9. The transfer is catalysed by a family of cysteine rich zinc finger protein acyltransferases that are characterized by a conserved D-H-H-C catalytic domain and are called DHHC proteins. 23 DHHC proteins are encoded in the humane genome. The DHHC proteins mediate the transfer in two steps. First, the DHHC protein is autoacylated. This happens through the reaction of palmityl-CoA with the cysteine residue in the catalytic domain of the DHHC protein forming an acyl-intermediate while simultaneously releasing CoA-SH. The intermediate is then directly transferred from the DHHC protein onto the substrate protein8,9. Thus, the molecular mass of the protein is increased by 238 Da for each palmitate moiety10. The abbreviated mechanism of S-palmitoylation is shown in figure 2.
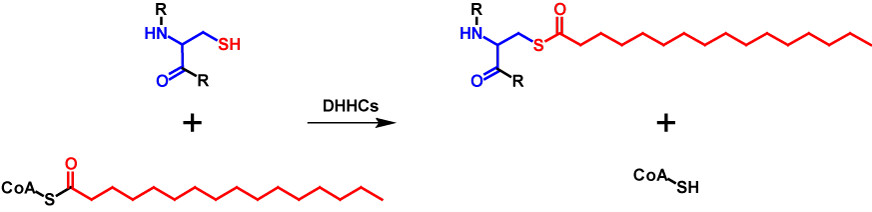
Sometimes S-palmitoylation refers to the covalent attachment of several medium- to long-chain fatty acids and not exclusively palmitate3,9. However, this is incorrect and should be referred to as S-acylation8. S-Palmitoylation influences localization, stability, interaction with effector proteins, enzyme activity, membrane trafficking and other parts of the cellular processes. For membrane proteins, the primary role of S-palmitoylation is to ease membrane attachment. This modification is present in all eukaryotic cells8,9. In contrast to N-myristoylation, S-palmitoylation has no conserved consensus sequence and is a reversible modification. The fatty acid moieties can be removed by serine hydrolases3,8,9.
References
- 1.Thinon E, Hang HC. Chemical reporters for exploring protein acylation. Biochemical Society transactions. 2015;43:253–261. doi:10.1042/BST20150004
- 2.Khoury GA, Baliban RC, Floudas CA. Proteome-wide post-translational modification statistics: frequency analysis and curation of the swiss-prot database. Scientific Reports. 2011;1:90. doi:10.1038/srep00090
- 3.Resh MD. Trafficking and signaling by fatty-acylated and prenylated proteins. Nature Chemical Biology. 2006;2:584–590. doi:10.1038/nchembio834
- 4.Farazi TA, Waksman G, Gordon JI. The Biology and Enzymology of ProteinN-Myristoylation . Journal of Biological Chemistry . 2001;276:39501–39504. http://www.jbc.org/content/276/43/39501.short
- 5.Wright MH, Heal WP, Mann DJ, Tate EW. Protein myristoylation in health and disease. Journal of Chemical Biology. 2010;3:19–35. doi:10.1007/s12154-009-0032-8
- 6.Legrand P, Rioux V. The Complex and Important Cellular and Metabolic Functions of Saturated Fatty Acids. Lipids. 2010;45:941–946. doi:10.1007/s11745-010-3444-x
- 7.Chen TF, Yoder JD, Hruby DE. Mass spectrometry analysis of synthetically myristoylated peptides. European journal of mass spectrometry (Chichester, England). 2004;10:501–508. doi:10.1255/ejms.652
- 8.De I, Sadhukhan S. Emerging Roles of DHHC-mediated Protein S-palmitoylation in Physiological and Pathophysiological Context. European Journal of Cell Biology. 2018;97:319–338. doi:https://doi.org/10.1016/j.ejcb.2018.03.005
- 9.Ko P-J, Dixon SJ. Protein palmitoylation and cancer. EMBO reports. 2018;19. doi:10.15252/embr.201846666
- 10.Rodenburg RNP, Snijder J, van de Waterbeemd M, et al. Stochastic palmitoylation of accessible cysteines in membrane proteins revealed by native mass spectrometry. Nature communications. 2017;8:1280. doi:10.1038/s41467-017-01461-z